One of the powerful library used for graph building activities is NetworkX. It is widely used in solving graph problems and network related queries. Lets have a look into NetworkX now.
In order to use it with python import it,
import networkx as nx
The following basic graph types are provided as Python classes:
Graph
This class implements an undirected graph. It ignores multiple edges between two nodes. It does allow
self-loop edges between a node and itself.
DiGraph
Directed graphs, that is, graphs with directed edges. Provides operations common to directed graphs, (a
subclass of Graph).
MultiGraph
A flexible graph class that allows multiple undirected edges between pairs of nodes. The additional
flexibility leads to some degradation in performance, though usually not significant.
MultiDiGraph
A directed version of a MultiGraph
g=nx.Graph() g.add_edge(1,3,weight=0.2) g.add_edge(3,2,weight=.3) nx.draw(g)
we can even make edges or not as per requirement, like the code below:
import matplotlib.pyplot as plt
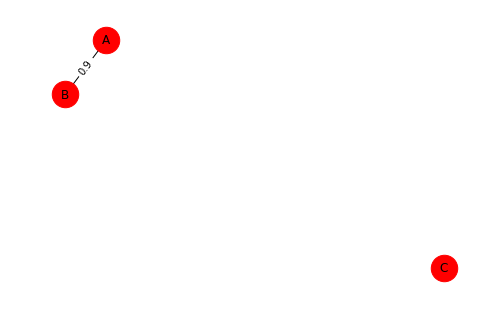
g2=nx.Graph()
g2.add_edge('A','B',weight=0.9)
g2.add_node('C')
#nx.draw(g2)
pos = nx.spring_layout(g2) # compute graph layout
nx.draw(g2, pos, node_size=700) # draw nodes and edges
nx.draw_networkx_labels(g2, pos)
labels = nx.get_edge_attributes(g2, 'weight')
nx.draw_networkx_edge_labels(g2, pos, edge_labels=labels)
plt.show(g2)
Now, if we go for a shortest path algorithm, networkx really rocks, lets take an example;
G=nx.Graph()
e = [('a', 'b', 0.3), ('b', 'c', 0.9), ('a', 'c', 0.5), ('c', 'd', 1.2)]
G.add_weighted_edges_from(e)
pos = nx.spring_layout(G)
nx.draw(G,pos,node_size=700)
nx.draw_networkx_labels(G, pos)
labels = nx.get_edge_attributes(G, 'weight')
nx.draw_networkx_edge_labels(G, pos, edge_labels=labels)
plt.show(G)
print(nx.dijkstra_path(G, 'a', 'd'))
['a', 'c', 'd']
We can even write few modular code to check the adjacency matrix and number of nodes relationship,
#Asignment 2 Comparision
for i in range(3):
GraphA=lstA[i]
GraphB=lstB[i]
Aad = nx.adjacency_matrix(GraphA)
Bad = nx.adjacency_matrix(GraphB)
count=i
count=count+1
print('-------------****************-------------')
print('Number of nodes: ',count*100)
for k in range(2,4):
print('An^k','n',count*100,'k',k)
A_nk=np.linalg.matrix_power(Aad.todense(), k)
#print(B_nk)
plt.plot(A_nk)
plt.show()
print('Bn^k','n',count*100,'k',k)
B_nk=np.linalg.matrix_power(Bad.todense(), k)
#print(B_nk)
plt.plot(B_nk)
plt.show()
print('------************------')
-------------****************------------- Number of nodes: 100 An^k n 100 k 2
Bn^k n 100 k 2


Having read this I thought it was extremely informative.
I appreciate you finding the time and energy to put this
information together. I once again find myself personally spending a
lot of time both reading and commenting. But so what, it
was still worthwhile!
Somebody necessarily assist to make critically articles I would state. This is the very first time I frequented your website page and up to now? I amazed with the research you made to make this actual post incredible. Magnificent activity!
Its like you read my mind! You appear to know
a lot about this, like you wrote the book in it or something.
I think that you could do with a few pics to drive the message home a little bit, but instead of that, this
is magnificent blog. An excellent read. I’ll certainly be back.